Deletion of retinoic acid-related orphan receptor gamma reduces body weight and hepatic lipids in mice by modulating the expression of lipid metabolism genes
Abstract
Aim: Retinoic acid-related orphan receptor γ (RORγ) functions as a ligand-dependent transcription factor and its loss has been shown to affect the circadian expression of lipid metabolism genes. However, its effect on body weight gain and hepatic lipids is not well understood. In this study, we investigated the impact of Rorγ gene deletion on changes in body weight and hepatic lipids.
Methods: Body weight and lipids were analyzed in the plasma and liver. Expression of lipid metabolism genes in the liver was evaluated in wild type and Rorγ knockout mice.
Results: We show that deletion of RORγ results in reduced body weight and fewer lipids in the liver. Analysis of gene expression showed that deletion of Rorγ resulted in an overall lower expression of genes and transcription factors involved in lipid biosynthesis. We observed a decrease in the gene expression of cholesterol biosynthesis, efflux, and esterification but an increase in bile acid synthesis. There was a decrease in fatty acid and triglycerides biosynthesis genes and an increase in the fatty acid uptake genes. The decrease in the expression of lipid biosynthesis genes was accompanied by the decrease in the sterol response element binding protein (Srebp) genes. We observed an increase in the expression of peroxisome proliferator-activated receptor alpha (Ppara) and a decrease in the expression of acetyl-CoA carboxylase 2 (Acc2) genes.
Conclusion: Our data suggest that RORγ regulates body weight and lipid metabolism genes and its modulation may be beneficial for the management of obesity and related lipid metabolic disorders.
Keywords
Introduction
Obesity is a major risk factor that leads to the development of other metabolic disorders such as hyperlipidemia and hepatic steatosis. Ectopic accumulation of fat in the liver causes insulin resistance by activating various cellular stress and inflammatory signaling pathways[1]. The liver plays an important role in lipid metabolism that involves several key proteins and transcription factors to maintain lipid homeostasis[2]. Various cellular regulators that include several transcription factors control biosynthesis, oxidation, uptake, and secretion of lipids in the liver[3]. Peroxisome proliferator-activated receptors (PPARs), which serve as hepatic lipid sensors, have been demonstrated to play a critical role in lipid metabolism by controlling the enzymes through various signaling events[4,5]. Peroxisome proliferator-activated receptor α (PPARα) is highly expressed in the liver and is primarily involved in the regulation of enzymes crucial for fatty acid oxidation. It also plays a role in regulating genes that control fatty acid elongation, desaturation and transport[6]. On the other hand, peroxisome proliferator-activated receptor γ (PPARγ), which is expressed at low levels in the liver[7], controls the upregulation of a subset of the lipogenic genes that are involved in lipid synthesis and accumulation[8]. Besides PPARs, the sterol response element binding proteins (SREBPs) that act as nutrient sensing transcription factors are also involved in the transcriptional control of the lipogenic gene expression[9]. Sterol regulatory-element binding protein 1c (SREBP-1c) is a transcription factor that primarily upregulates the transcription of genes involved in fatty acid synthesis. In contrast, sterol regulatory-element binding protein 2 (SREBP-2) controls the activation of genes responsible mainly for cholesterol synthesis and uptake, as opposed to fatty acid synthesis. However, sterol regulatory-element binding protein 1a (SREBP-1a) leads to the activation of genes involved in both pathways[10,11].
The transcriptional networks of orphan nuclear receptors govern the hepatic metabolic pathways[12]. The retinoic acid-related orphan receptor γ (RORγ), a member of the ROR subfamily of nuclear receptors, has been implicated in the control of a variety of physiological processes[13]. RORγ binds to specific ROR-responsive elements at its genomic targets to constitutively activate the gene expression in the absence of a ligand[14]. Global RORγ-deficient mice are born healthy and fertile but die within the first four months after birth due to high incidence of T-cell lymphomas in the thymus[15]. The physiological functions of RORγ, expressed in various peripheral tissues[16], are still poorly understood. RORγ has been shown to regulate the circadian expression of glucose and lipid metabolism genes[17,18]. Pharmacological inhibition of RORγ has been shown to reduce food intake and body weight gain[19]. However, effect of Rorγ gene deletion on body weight and hepatic lipids is not well studied. Therefore, in this study, we investigated the effect of Rorγ gene deletion on body weight and hepatic lipids using a knockout (KO) mouse model. We also studied the expression of downstream lipid metabolism genes that are regulated directly or indirectly by RORγ.
Methods
Materials
Infinity cholesterol (catalog # TR13421) and Infinity triglyceride (catalog # TR22421) reagent kits were purchased from Thermo Scientific (Middletown, VA). TRIzolTM (catalog #15596018) was purchased from Life Technologies (Carlsbad, CA). Omniscript RT kit (catalog #205113) was from Qiagen (Germantown, MD). qPCRTM core kit for SYBR Green I (catalog #10-SN10-05) was purchased from Eurogentec (San Diego, CA). All other chemicals and solvents were obtained from Fisher Scientific through its local distributor in the Kingdom of Saudi Arabia.
Animals
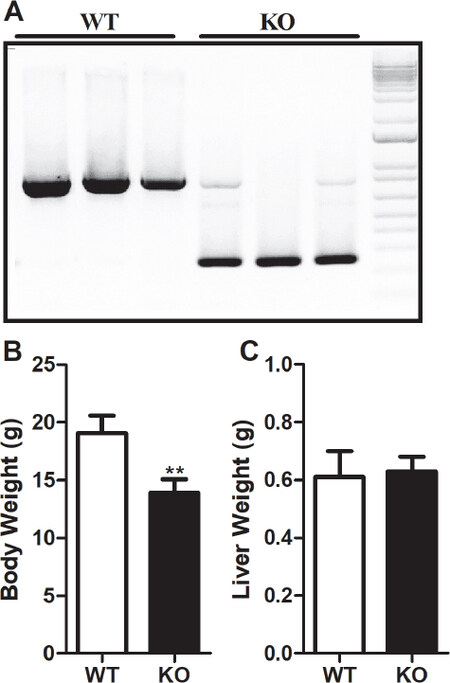
Heterozygous RORγ mice were intercrossed to obtain Rorγ-/- and wildtype (WT) littermate controls as described previously[20]. Genotyping was performed by PCR analysis of tail DNA using 5’-TGGTAGTGTCACTATCTGTGTCCC-3’ forward and 5’-CTTCAGAACTTATGTCAGGAACTCC-3’ reverse primers to generate an 850-bp WT product and a 250-bp KO product to identify the WT and Rorγ KO mice [Figure 1A]. For this study, we used eight-week-old chow-diet-fed WT and Rorγ KO male mice (n = 3) on C57BL/6J background. All experiments were approved by Institutional Animal Care and Use Committee at King Faisal Specialist Hospital and Research Center.
Figure 1. Effect of RORγ gene deletion on body and liver weights in mice. RORγ WT and KO mice (n = 3) were identified by genotyping the DNA isolated from tail tissues (A). Total body (B) and liver (C) weights of 8-week old chow diet fed WT and KO mice were measured. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. **P < 0.01. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ
Body weight and lipid analysis
Mice in WT and Rorγ KO groups were fasted overnight for 18 h, weighed using digital balance to measure total body weight and sacrificed to collect blood and livers. Liver weight of each animal was recorded. Blood was used to isolate plasma after centrifugation of the samples for 15 min at 2500 rpm using a tabletop centrifuge. Plasma and liver tissues were stored at -80 °C until further analysis. Hepatic lipids from WT and KO mice were extracted by Bligh and Dyer method[21]. Total cholesterol and triglyceride levels in the plasma and liver tissues were measured using commercially available kits from Thermo Scientific as described previously[22].
Analysis of gene expression by quantitative real time PCR
TRIzolTM was used to isolate total RNA from tissues as per the manufacturer’s instructions. cDNA was synthesized using Omniscript RT kit from good quality RNA preparations with the purity of RNA having A260/A280 ratio of more than 1.7. Quantitative PCR in each reaction was carried out in a total volume of 20 µL, consisting of 5 µL of 1:100 diluted first strand cDNA sample and 15 µL of SYBR Green I PCR master mix solution. The reaction mixture was incubated for 10 min at 95 °C followed by 40 cycles of 15 s incubations at 95 °C and 60 s at 60 °C in a QuantStudioTM 6 Flex Real-Time PCR (Applied Biosystems). Data were analyzed and presented as arbitrary units that were normalized to the expression of 18 sRNA, according to the manufacturer’s instructions. The primers were designed using Primer Express 3.0 (Applied Biosystems) and are shown in Table 1.
List of primers used for the quantification of lipid metabolism genes
| Gene | Forward primer | Reverse primer |
|---|---|---|
| Cholesterol metabolism genes | ||
| Hmgr | AGCTTGCCCGAATTGTATGT | TCTGTTGTGAACCATGTGAC |
| Hmgs | TGGCTATAAAGCTGCGGAGG | GGTGAAAGAGCCAAAGGGGA |
| Abca1 | GGACATGCACAAGGTCCTGA | CAGAAAATCCTGGAGCTTCA |
| Abcg1 | GCTGTGCGTTTTGTGCTGTT | TGCAGCTCCAATCAGTAGGC |
| Abcg5 | AGGGCCTCACATCAACAGAG | GCTGACGCTGTAGGACACAT |
| Abcg8 | AGTGGTCAGTCCAACACTCT | GAGACCTCCAGGGTATCTTG |
| Acat1 | CCAATGCCAGCACACTGAAC | TCTACGGCAGCATCAGCAAA |
| Acat2 | TGTTGAAAGGTGGGCAGCAA | CAGGTAACATCCCATCCCGT |
| Cyp7a | CAACGGGTTGATTCCATACC | ATTTCCCCATCAGTTTGCAG |
| Fatty acid metabolism genes | ||
| Cd36 | TGCACCACATATCTACCAAA | TTGTAACCCCACAAGAGTTC |
| Fabp1 | CAATAGGTCTGCCCGAGGAC | CAGGGTGAACTCATTGCGGA |
| Fas | GGGTTCTAGCCAGCAGAGTC | TCAGCCACTTGAGTGTCCTC |
| Acc1a | GAGGAAGTTGGCTATCCAGT | CTTGAACCTGTCTGAAGAGG |
| Acc2 | TGTCCCAGGAGGCTGCATTG | TGTGCAGGTCCAGTTTCTTG |
| Scd1 | CCGGAGACCCCTTAGATCGA | TAGCCTGTAAAAGATTTCTG |
| Triglycerides metabolism genes | ||
| Dgat1 | GTTCAGCTCAGACAGTGGTT | TCAGCATCACCACACACCAA |
| Dgat2 | AGCTGCAGGTCATCTCAGTA | CTGCAGGCCACTCCTAGCAC |
| Mgat2 | GAGCAAAGCCCGTGTGTAGA | AAGGTCTGTAACCTGCGCTC |
| Mttp | CACACAACTGGCCTCTCATTAAAT | TGCCCCCATCAAGAAACACT |
| Vldlr | GCCATATGAGAACATGCCGC | AGGACACGGGGATACACTGA |
| Ldlr | TGACTCAGACGAACAAGGCT | ATCTAGGCAATCTCGGTCTC |
| Plin1 | TGCTGCACGTGGAGAGTAAG | AGCAGGGTTGGGCCCTTGTT |
| Lpl | TTTGGCTCCAGAGTTTGACC | TGTGTCTTCAGGGGTCCTTA |
| Apoc2 | CATGGGGTCTCGGTTCTTCC | CTTAAGAGGGAGCCCTGCAC |
| Lipid metabolism transcription factor genes | ||
| Srebp1a | CCATGGACGAGCTGGCCTTC | AGTTGGCACCTGGGCTGCTG |
| Srebp1c | GGAAGCTGTCGGGGTAGCGT | CATGTCTTCAAATGTGCAAT |
| Srebp2 | GCGTTCTGGAGACCATGGAG | GAGCTACAAAGTTGCTCTGA |
| Ppara | GCGTACGGCAATGGCTTTAT | GAACGGCTTCCTCAGGTTCT |
| Pparg | ATGGTGCCTTCGCTGATGCA | TGGCATCTCTGTGTCAACCA |
| Housekeeping gene | ||
| 18 sRna | GTAACCCGTTGAACCCCATT | CCATCCAATCGGTAGTAGCG |
Statistical analysis
Data are presented as mean ± SD Statistical significance (P < 0.05) was determined using Student’s t test (GraphPad Prism 5).
Results
Effect of Rorγ gene deletion on body and liver weights
Age matched eight-week-old male Rorγ KO mice and littermate controls on chow diet were used to assess the effect of Rorγ gene deletion on total body and liver weights. Rorγ gene deletion resulted in a significant decrease of 27% in total body weight compared to WT littermate control mice [Figure 1B]. However, no change was seen in the weight of livers in these mice [Figure 1C]. These results suggest that RORγ is important in maintaining the body weight in mice and its deficiency leads to reduced body weight gain.
RORγ KO mice decreases hepatic but not plasma lipids
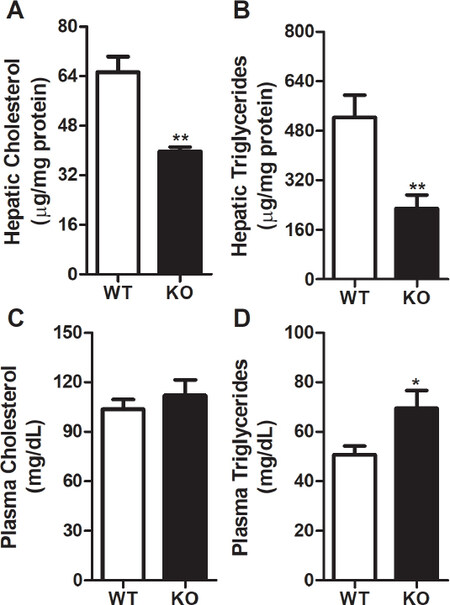
Next, we investigated whether change in body weight after Rorγ gene deletion also affects liver and plasma lipids. Our data indicate that there were significant decreases of 39% and 56% in the levels of liver cholesterol [Figure 2A] and triglycerides [Figure 2B], respectively, in Rorγ KO mice compared to WT littermates. However, we did not see any significant change in plasma cholesterol levels between WT and Rorγ KO mice [Figure 2C]. Contrary to decreased hepatic lipid levels, we observed an increase of ~37% in the levels of plasma triglycerides in KO mice compared to WT mice [Figure 2D]. These data suggest that Rorγ gene deletion affects only liver and not plasma cholesterol. On the other hand, both liver and plasma triglycerides are altered after the ablation of Rorγ gene.
Figure 2. RORγ gene deletion reduces hepatic lipids and increases plasma triglycerides. Lipids from WT and KO mice livers (n = 3) were extracted by Bligh and Dyer method to measure cholesterol (A) and triglycerides (B). Total cholesterol (C) and triglycerides (D) were also measured in the plasma of these mice. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. **P < 0.01. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ
Ablation of Rorγ activity affects expression of genes involved in lipid metabolism
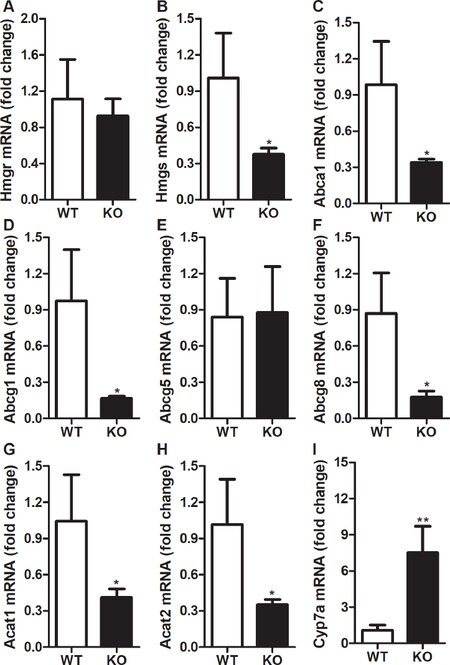
To gain better understanding of how Rorγ activity regulates body weight and lipids, we looked at the changes in the expression of lipid metabolism genes after the deletion of Rorγ gene. We started by analyzing the changes in the genes that regulate cholesterol homeostasis in the liver. Coordinated expression of several genes regulate cellular cholesterol metabolism[23,24]. We looked at the expression of cholesterol biosynthesis genes such as hydroxymethylglutaryl-CoA reductase (Hmgr) and hydroxymethylglutaryl-CoA synthase (Hmgs). The expression of rate-limiting enzyme gene, Hmgr, was not affected by the deletion of Rorγ in the liver [Figure 3A]. However, we observed a decrease of 63% in the expression of Hmgs in the livers of Rorγ KO mice compared to WT littermates [Figure 3B]. Next, we looked at the expression of genes involved in the efflux of cholesterol in the liver. There was a decrease of 65%-83% in the expression of ATP-binding cassette transporter A1 (Abca1), G1 (Abcg1), and G8 (Abcg8) but not G5 (Abcg5) [Figure 3C-F]. Besides the decrease in cholesterol biosynthesis and efflux genes, we also observed a 61%-65% decrease in the expression of acetyl-CoA acetyltransferase 1 (Acat1) and acetyl-CoA acetyltransferase 2 (Acat2) genes that are involved in the esterification of cholesterol for storage and lipoprotein secretion [Figure 3G and H]. On the other hand, expression of cholesterol 7alpha-hydroxylase (Cyp7a) involved in the biosynthesis of bile acids was increased by ~6 folds in the livers of Rorγ KO mice [Figure 3I]. Taken together, these results demonstrate that ablation of Rorγ activity results in decreased expression of genes involved in cholesterol biosynthesis, efflux, and storage but increased expression of genes involved in bile acid biosynthesis.
Figure 3. Changes in hepatic expression of cholesterol metabolism genes in RORγ knockout mice. Total mRNA from the livers of WT and KO mice were used to determine the expression of cholesterol metabolism genes. Relative changes in the mRNA expression were calculated based on the expression of 18 sRNA. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. *P < 0.05 and **P < 0.01. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ; Hmgr: hydroxymethylglutaryl-CoA reductase; Hmgs: hydroxymethylglutaryl-CoA synthase; Abca1: ATP-binding cassette transporter A1; Abcg1: ATP-binding cassette transporter G1; Abcg5: ATP-binding cassette transporter G5; Abcg8: ATP-binding cassette transporter G8; Acat1: acetyl-CoA acetyltransferase 1; Acat2: acetyl-CoA acetyltransferase 2; Cyp7a: cholesterol 7alpha-hydroxylase
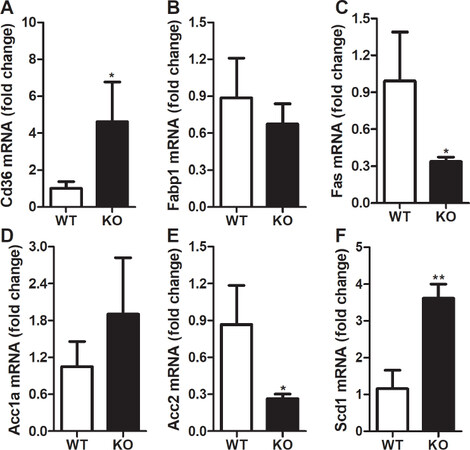
We then looked at the changes in the expression of fatty acid metabolism genes. Uptake of long-chain fatty acids by the liver is dependent on fatty acid transporters such as cluster of differentiation 36 (Cd36). There was a ~4.5-fold increase in the expression of Cd36 in the livers of KO mice compared to WT mice [Figure 4A]. However, we did not see any change in the expression of fatty acid binding protein 1 (Fabp1) in the livers of Rorγ KO mice [Figure 4B]. Next, we looked at the expression of fatty acid synthesis genes and found that deletion of Rorγ decreased the expression of fatty acid synthase (Fas) and acetyl-CoA carboxylase 2 (Acc2) by 66% [Figure 4C] and 70% [Figure 4E], respectively, but had no significant effect on the expression of acetyl-CoA carboxylase 1a (Acc1a) [Figure 4D]. Interestingly, expression of stearoyl-coenzyme A desaturase-1 (Scd1), which catalyzes the conversion of saturated fatty acids to monounsaturated fatty acids, was increased by ~3 folds in the livers of Rorγ KO mice [Figure 4F]. These results suggest that deletion of Rorγ increases the expression of genes involved in fatty acids uptake and desaturation but decreases the expression of genes involved in their biosynthesis.
Figure 4. Changes in hepatic expression of fatty acid metabolism genes in RORγ knockout mice. Total mRNA from the livers of WT and KO mice were used to determine the expression of fatty acid metabolism genes. Relative changes in the mRNA expression were calculated based on the expression of 18 sRNA. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. *P < 0.05 and **P < 0.01. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ; Cd36: cluster of differentiation 36; Fabp1: fatty acid binding protein 1; Fas: fatty acid synthase; Acc1a: acetyl-CoA carboxylase 1a; Acc2: acetyl-CoA carboxylase 2; Scd1: stearoyl-coenzyme A desaturase-1
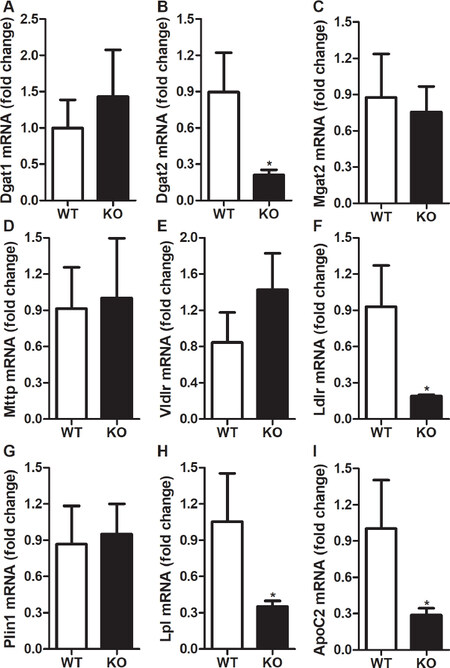
We further investigated the effect of Rorγ gene deletion on the expression of genes involved in triglycerides metabolism. There was a significant reduction of 77% in the expression of diacylglycerol O-acyltransferase 2 (Dgat2) gene [Figure 5B] in contrast to a small but not significant increase in the expression of Dgat1 gene [Figure 5A], and no change in the expression of acyl CoA: monoacylglycerol acyltransferase 2 (Mgat2) gene [Figure 5C] in the livers of Rorγ KO mice as compared to WT mice. Dgat2 is mainly expressed and involved in the synthesis of bulk of triglycerides in mouse liver in contrast to Dgat1 and Mgat2, which are highly expressed in the small intestine[25,26]. Our data also show that deletion of Rorγ gene did not affect the expression of microsomal triglyceride transfer protein (Mttp) gene, a key protein involved in the assembly and secretion of lipid rich lipoproteins[27,28][Figure 5D]. Very low-density lipoprotein receptor (Vldlr) and low-density lipoprotein receptor (Ldlr) are important proteins that regulate the uptake of lipid rich lipoproteins from the circulation. There was no significant difference in the expression of Vldlr between the Rorγ KO and WT mice [Figure 5E] but we observed a 79% decrease in Ldlr gene expression in the livers of KO mice compared to WT mice [Figure 5F]. Furthermore, there was no difference in the expression of lipid droplet protein, perilipin 1 (Plin1), in the livers of Rorγ KO and WT mice [Figure 5G]. To understand the reasons for the increase in plasma triglycerides, we looked at the expression of lipoprotein lipase (Lpl) and apolipoprotein C2 (ApoC2), which are involved in the lipolysis of lipids in the circulation. Expression of both these genes was reduced by 67%-71% in Rorγ KO mice [Figure 5H, I]. These results suggest that expression of genes involved in triglycerides synthesis, uptake, and lipolysis is decreased in Rorγ KO mice leading to decreased triglycerides accumulation in the liver and increased triglycerides in the plasma.
Figure 5. Changes in hepatic expression of triglycerides metabolism genes in RORγ knockout mice. Total mRNA from the livers of WT and KO mice were used to determine the expression of triglycerides metabolism genes. Relative changes in the mRNA expression were calculated based on the expression of 18 sRNA. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. *P < 0.05. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ; Dgat1: diacylglycerol O-acyltransferase 1; Dgat2: diacylglycerol O-acyltransferase 2; Mgat2: acyl CoA:monoacylglycerol acyltransferase 2; Mttp: microsomal triglyceride transfer protein; Vldlr: very low-density lipoprotein receptor; Ldlr: low-density lipoprotein receptor; Plin1: perilipin 1; Lpl: lipoprotein lipase; Apoc2: apolipoprotein C2
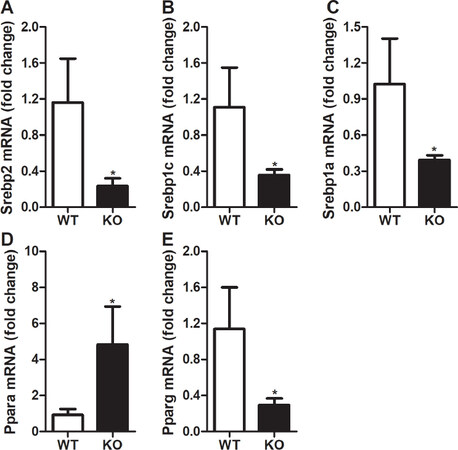
To address whether alterations in cellular lipids is due to the changes in the expression of nuclear transcription factors, we looked at the expression of these genes in the liver. Our data indicate that the deletion of Rorγ reduced Srebp2 gene expression by 80% in the livers [Figure 6A]. These data are consistent with the lower hepatic cholesterol levels [Figure 2A] and decreased expression of downstream genes involved in cholesterol biosynthesis pathway [Figure 3]. Our results show that ablation of Rorγ activity in the liver leads to a 68% decrease in the expression of Srebp1c gene [Figure 6B], which is consistent with the reduced hepatic triglycerides [Figure 2B] and lower expression of the lipid synthesis genes [Figures 4 and 5]. Furthermore, we also noticed that expression of Srebp1a gene was significantly reduced by 62% in the livers of Rorγ KO mice compared to WT littermates [Figure 6C]. Next, we looked at the expression of Ppara and Pparg genes in the livers of Rorγ KO and WT mice. There was an around five-fold increase in the expression of Ppara gene, which is mainly involved in fatty acid oxidation [Figure 6D]. However, we observed a decrease of 74% in the expression of Pparg gene [Figure 6E]. These data suggest that expression of transcription factors that regulate lipid metabolism is modulated by RORγ.
Figure 6. Changes in hepatic expression of lipid metabolism transcription factor genes in RORγ knockout mice. Total mRNA from the livers of WT and KO mice were used to determine the expression of transcription factor genes. Relative changes in the mRNA expression were calculated based on the expression of 18 sRNA. Values were plotted as mean ± SD. P values were calculated using two-tailed Student’s t test. *P < 0.05. WT: wild type; KO: knockout; RORγ: retinoic acid-related orphan receptor γ; Srebp2: sterol regulatory-element binding protein 2; Srebp1c: sterol regulatory-element binding protein 1c; Srebp1a: sterol regulatory-element binding protein 1a; Pparα: peroxisome proliferator-activated receptor α; Pparγ : peroxisome proliferator-activated receptor γ
Discussion
Our results identify a critical role of RORγ in lipid metabolism. Here, we show that ablation of Rorγ gene decreases the body weight and reduces the accumulation of lipids in the liver. Interestingly, we did not see any significant change in the liver weight between the WT and Rorγ KO mice [Figure 1C]; however, we noticed a visible decrease in the content of abdominal fat mass in Rorγ KO mice (data not shown). This decrease in body weight and fat mass may be due to either reduced food intake or increased energy expenditure. It is known that activation of PPARγ by agonists promote weight gain and fat accumulation mainly due to increased adipocyte differentiation and lipid storage[29,30]. Our data indicate that Rorγ gene deletion results in lower expression of Pparg[Figure 6E], which may explain the decrease in body weight and fat mass. This finding is consistent with the repression of Rorγ activity by inverse agonist that lead to a reduction in fat mass and body weight in obese diabetic mice[19].
A decrease in body weight and fat mass in Rorγ KO mice suggests that these mice do not store enough lipids in the adipocytes. Inability of adipocytes to store the excess fatty acids coming from the diet suggest that these fatty acids may be taken up by the liver for either oxidation or synthesis of lipids for storage in hepatic or extrahepatic tissues such as muscles. A significant increase in the hepatic expression of Cd36 gene responsible for the uptake of long-chain fatty acids suggests that the livers from Rorγ KO mice may be taking up more fatty acids from the circulation [Figure 4A]. Takeda et al.[17] also showed that expression of Cd36 gene was upregulated at all the times during diurnal oscillations in the livers of Rorγ KO mice. Since both cholesterol and triglycerides levels were reduced in the livers of Rorγ KO mice [Figure 2A and B], we speculate that most of the fatty acids taken up by these mice are not utilized for lipid biosynthesis and storage. This was further supported by our observation that the expression of genes involved in lipid biosynthesis was lower in the livers from Rorγ KO mice [Figure 5B]. It is interesting to note that there was a reduction in the expression of Acc2 in the livers of Rorγ KO mice. This enzyme is involved in the synthesis of malonyl-coA that plays an important role in regulating the oxidation of fatty acids in the mitochondria by inhibiting the carnitine/palmitoyl shuttle system[31] in contrast to malonyl-coA that is generated by the ACC1 and utilized by Fas for the synthesis of fatty acids in the cytosol[32]. Therefore, lower levels of Acc2 gene expression suggests that the oxidation of fatty acids may be enhanced in the livers of Rorγ KO mice, which is consistent with the inhibition of Rorγ activity by inverse agonist that leads to enhanced fatty acid oxidation[19]. Hence, besides reduced lipid biosynthesis, increased fatty acid oxidation may also contribute to reduced accumulation of triglycerides in the livers of Rorγ KO mice.
Contrary to lower hepatic lipids, we observed an increase in the levels of triglycerides in the plasma of Rorγ KO mice. On the other hand, Takeda et al.[17] reported a decrease in the levels of plasma lipids in Rorγ KO mice. The discrepancy in results may be due to the feeding of high fat diet for six weeks to these mice, which may affect the metabolism of lipids in the plasma. It is likely that Rorγ gene ablation affects the uptake or secretion of these lipids from the liver. We did not observe any changes in the expression of Mttp, a gene responsible for the secretion of lipid rich lipoproteins, suggesting that secretion of lipids by the liver may not be affected by Rorγ gene deletion. However, our data indicate that Rorγ gene deletion decrease the levels of Ldlr gene expression implicating that uptake of lipid rich lipoproteins may be reduced in the KO mice leading to increase in the levels of plasma triglycerides. It is possible that the increase in plasma triglycerides may also be due to reduced activity of lipoprotein lipase. In fact, our data show that expression of Lpl and ApoC2 genes were reduced in the livers of Rorγ KO mice. ApoC2 acts as an activator of lipoprotein lipase[33] and its reduction in the circulation may lead to reduced lipolysis, thereby causing increased plasma triglycerides.
RORγ functions as a transcriptional mediator that has been shown to regulate the rhythmic expression of certain metabolic genes downstream of clock machinery[17,18]. Regulation of these genes is complex and may involve other nuclear receptors or transcriptional factors downstream of Rorγ to modulate lipid metabolism directly or indirectly[17]. There is also a possibility that changes in these transcriptional factors may be secondary to the changes in the homeostasis of lipids in the cells[34]. Expression of several lipid metabolism genes is regulated by various SREBPs that sense the changes in lipid levels in the cells. Stimulation of SREBP-2 regulates cholesterol homeostasis by activating the transcription of genes involved cholesterol metabolism[35]. On the other hand, stimulation of SREBP-1c increases lipogenesis by increasing the expression of fatty acid metabolism genes[36]. SREBP-1a has been shown to be an important regulator of both cholesterol and lipid synthesis genes[37]. Our results suggest that deletion of Rorγ decreases the expression of Srebp1a, Srebp1c, and Srebp2 in the liver [Figure 6]. Changes in the expression of these transcription factors decreased the overall biosynthesis of cholesterol, fatty acids, and triglycerides in the liver cells. Our data also indicate that deletion of Rorγ may increase the oxidation of fatty acids due to increased expression of Ppara in the liver.
We observed that expression of genes involved in biosynthesis, efflux, and esterification of cholesterol was down-regulated in Rorγ KO mice [Figure 3]. This may be the reason for less cholesterol levels in the livers of these mice [Figure 2A]. Interestingly, we noticed a significant increase in the expression of Cyp7a gene, which is responsible for the biosynthesis of bile acids in the liver. Although we do not know the exact mechanism of how hepatic bile acid signaling was impaired in Rorγ KO, we speculate that a larger portion of the hepatic cholesterol pool may be directed towards bile acid biosynthesis in the absence of RORγ. Takeda et al.[17] also observed an increase in the expression of Cyp7a gene in Rorγ KO mice fed a chow diet. However, when these mice were fed a high fat diet for eight weeks, the levels of their liver, serum, and fecal bile acid pools were reduced significantly. It is possible that the high fat diet is repressing the expression of some other genes involved in the bile acid biosynthesis pathway.
In this study, we demonstrated that RORγ plays a significant role in the regulation of the expression of lipid metabolism genes that are involved in lipid biosynthesis, storage, and fatty acid oxidation. Our data also suggest that a cross-talk may occur between various transcription factors to maintain overall lipid homeostasis in the body. Previously, it has been shown that hepatic circadian clock plays a critical role in regulating major aspects of lipid metabolism through RORγ activity[17]. In a different experimental setting, we expanded the investigation to look at the changes in several lipid metabolism genes that were not studied previously. We also looked at the expression of different transcription factors to study the cross-talk between RORγ and these transcription factors to regulate lipid homeostasis. Most of the findings in this current study are consistent and added substantive findings to the previously published reports[17,19].
In summary, we show here that ablation of RORγ activity reduce lipid biosynthesis but enhance fatty acid oxidation by modulating the expression of several lipid metabolism genes. However, this study has some limitations because global RORγ-deficient mice are born healthy and fertile but die within the first four months after birth due to high incidence of T-cell lymphomas in the thymus[15]. Therefore, further studies are warranted to show the effect of RORγ deficiency on lipid metabolism in liver-specific KO mice to avoid any complications or confounding effects due to T-cell lymphomas. We will extend the studies of Takeda et al.[17] to investigate role of RORγ on lipid metabolism genes in chow-fed mice that were not studied previously. We will also look at the expression of different transcription factors to study the cross-talk between these factors and RORg to regulate lipid homeostasis. Furthermore, these mice may also be a good model to study the effect of liver-specific Rorγ gene deletion on diet-induced obesity. In conclusion, it may be beneficial to target RORγ activity for the management of obesity and related lipid metabolic disorders.
Declarations
Authors’ contributionsConducted experiments: Jahangir Z, Veluru D, Otaibi AA, Mubarak SA, Subie BA, Alghanem AF
Contributed to the conception and design of the study: Iqbal J, Hawwari A, Bakillah A, Qarni AA
Performed data analysis and interpretation: Iqbal J, Bakillah A
Wrote the paper: Iqbal J, Bakillah A, Hawwari A
Availability of data and materialsAll data and materials and mice are publicly available.
Financial support and sponsorshipThis research was supported by National Science, Technology and Innovation Plan’s strategies technologies program in Saudi Arabia (KACST: 10-BIO950-20) and an internal fund (RAC # 2083003) by KFSH&RC to A.H. This work was also partially supported by the intramural grants RA16/024/A and RA17/013/A to Iqbal J and RA17/014/A to Alghanem AF from KAIMRC. Jahangir Z worked on this study as a high school summer trainee at KAIMRC.
Conflicts of interestAll authors declare that there are no conflicts of interest.
Ethical approval and consent to participateAll experiments were approved by Institutional Animal Care and Use Committee at King Faisal Specialist Hospital and Research Center.
Consent for publicationNot applicable.
Copyright© The Author(s) 2019.
REFERENCES
1. Rutkowski JM, Stern JH, Scherer PE. The cell biology of fat expansion. J Cell Biol 2015;208:501-12.
2. Van den BG. The role of the liver in metabolic homeostasis: implications for inborn errors of metabolism. J Inherit Metab Dis 1991;14:407-20.
3. Karagianni P, Talianidis I. Transcription factor networks regulating hepatic fatty acid metabolism. Biochim Biophys Acta 2015;1851:2-8.
4. Guo Y, Jolly RA, Halstead BW, Baker TK, Stutz JP, et al. Underlying mechanisms of pharmacology and toxicity of a novel PPAR agonist revealed using rodent and canine hepatocytes. Toxicol Sci 2007;96:294-309.
5. Schoonjans K, Watanabe M, Suzuki H, Mahfoudi A, Krey G, et al. Induction of the acyl-coenzyme A synthetase gene by fibrates and fatty acids is mediated by a peroxisome proliferator response element in the C promoter. J Biol Chem 199518;270:19269-76.
7. Semple RK, Chatterjee VK, O’Rahilly S. PPAR gamma and human metabolic disease. J Clin Invest 2006;116:581-9.
8. Lee YJ, Ko EH, Kim JE, Kim E, Lee H, et al. Nuclear receptor PPARgamma-regulated monoacylglycerol O-acyltransferase 1 (MGAT1) expression is responsible for the lipid accumulation in diet-induced hepatic steatosis. Proc Natl Acad Sci U S A 2012;109:13656-61.
9. Wong RH, Sul HS. Insulin signaling in fatty acid and fat synthesis: a transcriptional perspective. Curr Opin Pharmacol 2010;10:684-91.
10. Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest 2002;109:1125-31.
11. Horton JD, Shah NA, Warrington JA, Anderson NN, Park SW, et al. Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc Natl Acad Sci U S A 200314;100:12027-32.
12. Chawla A, Boisvert WA, Lee CH, Laffitte BA, Barak Y, et al. A PPAR gamma-LXR-ABCA1 pathway in macrophages is involved in cholesterol efflux and atherogenesis. Mol Cell 2001;7:161-71.
13. Solt LA, Burris TP. Action of RORs and their ligands in (patho) physiology. Trends Endocrinol Metab 2012;23:619-27.
14. Medvedev A, Yan ZH, Hirose T, Giguere V, Jetten AM. Cloning of a cDNA encoding the murine orphan receptor RZR/ROR gamma and characterization of its response element. Gene 1996;181:199-206.
15. Ueda E, Kurebayashi S, Sakaue M, Backlund M, Koller B, et al. High incidence of T-cell lymphomas in mice deficient in the retinoid-related orphan receptor RORgamma. Cancer Res 2002;62:901-9.
16. Jetten AM. Retinoid-related orphan receptors (RORs): critical roles in development, immunity, circadian rhythm, and cellular metabolism. Nucl Recept Signal 2009;7:e003.
17. Takeda Y, Kang HS, Lih FB, Jiang H, Blaner WS, et al. Retinoid acid-related orphan receptor gamma, RORgamma, participates in diurnal transcriptional regulation of lipid metabolic genes. Nucleic Acids Res 2014;42:10448-59.
18. Jetten AM, Kang HS, Takeda Y. Retinoic acid-related orphan receptors alpha and gamma: key regulators of lipid/glucose metabolism, inflammation, and insulin sensitivity. Front Endocrinol (Lausanne) 2013;4:1.
19. Chang MR, He Y, Khan TM, Kuruvilla DS, Garcia-Ordonez R, et al. Antiobesity effect of a small molecule repressor of RORgamma. Mol Pharmacol 2015;88:48-56.
20. Kurebayashi S, Ueda E, Sakaue M, Patel DD, Medvedev A, et al. Retinoid-related orphan receptor gamma (RORgamma) is essential for lymphoid organogenesis and controls apoptosis during thymopoiesis. Proc Natl Acad Sci U S A 2000;97:10132-7.
21. Bligh EG, Dyer WJ. A rapid method of total lipid extraction and purification. Can J Biochem Physiol 1959;37:911-7.
22. Iqbal J, Rudel LL, Hussain MM. Microsomal triglyceride transfer protein enhances cellular cholesteryl esterification by relieving product inhibition. J Biol Chem 2008;283:19967-80.
23. Rotllan N, Fernandez-Hernando C. MicroRNA Regulation of Cholesterol Metabolism. Cholesterol 2012;2012:847849.
25. Yen CL, Farese RV Jr. MGAT2, a monoacylglycerol acyltransferase expressed in the small intestine. J Biol Chem 2003;278:18532-7.
26. McLaren DG, Han S, Murphy BA, Wilsie L, Stout SJ, et al. DGAT2 inhibition alters aspects of triglyceride metabolism in rodents but not in non-human primates. Cell Metab 2018;27:1236-48.
27. Hussain MM, Shi J, Dreizen P. Microsomal triglyceride transfer protein and its role in apoB-lipoprotein assembly. J Lipid Res 2003;44:22-32.
28. Hussain MM, Iqbal J, Anwar K, Rava P, Dai K. Microsomal triglyceride transfer protein: a multifunctional protein. Front Biosci 2003;8:s500-6.
29. Kadowaki T, Hara K, Kubota N, Tobe K, Terauchi Y, et al. The role of PPARgamma in high-fat diet-induced obesity and insulin resistance. J Diabetes Complications 2002;16:41-5.
30. Rosen ED, Sarraf P, Troy AE, Bradwin G, Moore K, et al. PPAR gamma is required for the differentiation of adipose tissue in vivo and in vitro. Mol Cell 1999;4:611-7.
31. McGarry JD, Mannaerts GP, Foster DW. A possible role for malonyl-CoA in the regulation of hepatic fatty acid oxidation and ketogenesis. J Clin Invest 1977;60:265-70.
32. Ruderman NB, Saha AK, Kraegen EW. Minireview: malonyl CoA, AMP-activated protein kinase, and adiposity. Endocrinology 2003;144:5166-71.
33. Fojo SS, Brewer HB. Hypertriglyceridaemia due to genetic defects in lipoprotein lipase and apolipoprotein C-II. J Intern Med 1992;231:669-77.
34. Jump DB, Tripathy S, Depner CM. Fatty acid-regulated transcription factors in the liver. Annu Rev Nutr 2013;33:249-69.
35. Brown MS, Goldstein JL. The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 1997;89:331-40.
36. Stoeckman AK, Towle HC. The role of SREBP-1c in nutritional regulation of lipogenic enzyme gene expression. J Biol Chem 2002;277:27029-35.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Iqbal J, Jahangir Z, Veluru D, Otaibi AA, Mubarak SA, Subie BA, Alghanem AF, Qarni AA, Bakillah A, Hawwari A. Deletion of retinoic acid-related orphan receptor gamma reduces body weight and hepatic lipids in mice by modulating the expression of lipid metabolism genes. Vessel Plus 2019;3:40. http://dx.doi.org/10.20517/2574-1209.2019.28
AMA Style
Iqbal J, Jahangir Z, Veluru D, Otaibi AA, Mubarak SA, Subie BA, Alghanem AF, Qarni AA, Bakillah A, Hawwari A. Deletion of retinoic acid-related orphan receptor gamma reduces body weight and hepatic lipids in mice by modulating the expression of lipid metabolism genes. Vessel Plus. 2019; 3: 40. http://dx.doi.org/10.20517/2574-1209.2019.28
Chicago/Turabian Style
Iqbal, Jahangir, Zainab Jahangir, Divya Veluru, Abeer Al Otaibi, Sindiyan A. Mubarak, Badr Al Subie, Ahmad F. Alghanem, Ali Al Qarni, Ahmed Bakillah, Abbas Hawwari. 2019. "Deletion of retinoic acid-related orphan receptor gamma reduces body weight and hepatic lipids in mice by modulating the expression of lipid metabolism genes" Vessel Plus. 3: 40. http://dx.doi.org/10.20517/2574-1209.2019.28
ACS Style
Iqbal, J.; Jahangir Z.; Veluru D.; Otaibi AA.; Mubarak SA.; Subie BA.; Alghanem AF.; Qarni AA.; Bakillah A.; Hawwari A. Deletion of retinoic acid-related orphan receptor gamma reduces body weight and hepatic lipids in mice by modulating the expression of lipid metabolism genes. Vessel Plus. 2019, 3, 40. http://dx.doi.org/10.20517/2574-1209.2019.28
About This Article
Copyright
Data & Comments
Data

 Cite This Article 7 clicks
Cite This Article 7 clicks










Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.